- 学院2023年研究生招生复试录...04/12
- 威人斯所有官方网站召开研究生导师培训会06/02
- 威人斯所有官方网站2022年研究生复试工...04/16
- 学院组织研究生赴张家口开展...06/23
- 2021届农村发展领域研究生毕...05/13
- 2021届校级优秀毕业研究生04/20
- 2020-2021学年度校级三好学生04/20
- 王述芬博士作国贸专业论文写作…11/09
- 国贸教学部召开国际贸易从业技…11/09
- 财务管理专业邀请中兴财光华会…11/08
- 威人斯所有官方网站邀请中国农业大学廖媛…04/17
- 威人斯所有官方网站举办重点项目申报辅导…04/17
- 学院举办系列科研讲座07/07
- 威人斯所有官方网站举办“省社科基金申报规范…04/28
- 威人斯所有官方网站获得第二届校园心理剧大…12/15
- 威人斯所有官方网站举办“我把旧貌换新颜”…12/04
- 威人斯所有官方网站志愿者赴白塔岭新区开展…12/04
- 威人斯所有官方网站志愿者赴晨旭融合幼教园…12/04
- 威人斯所有官方网站组织开展第四次团课11/30
- 学院心理剧比赛圆满结束11/24
- 学院团委组织开展“清洁幼儿…11/20
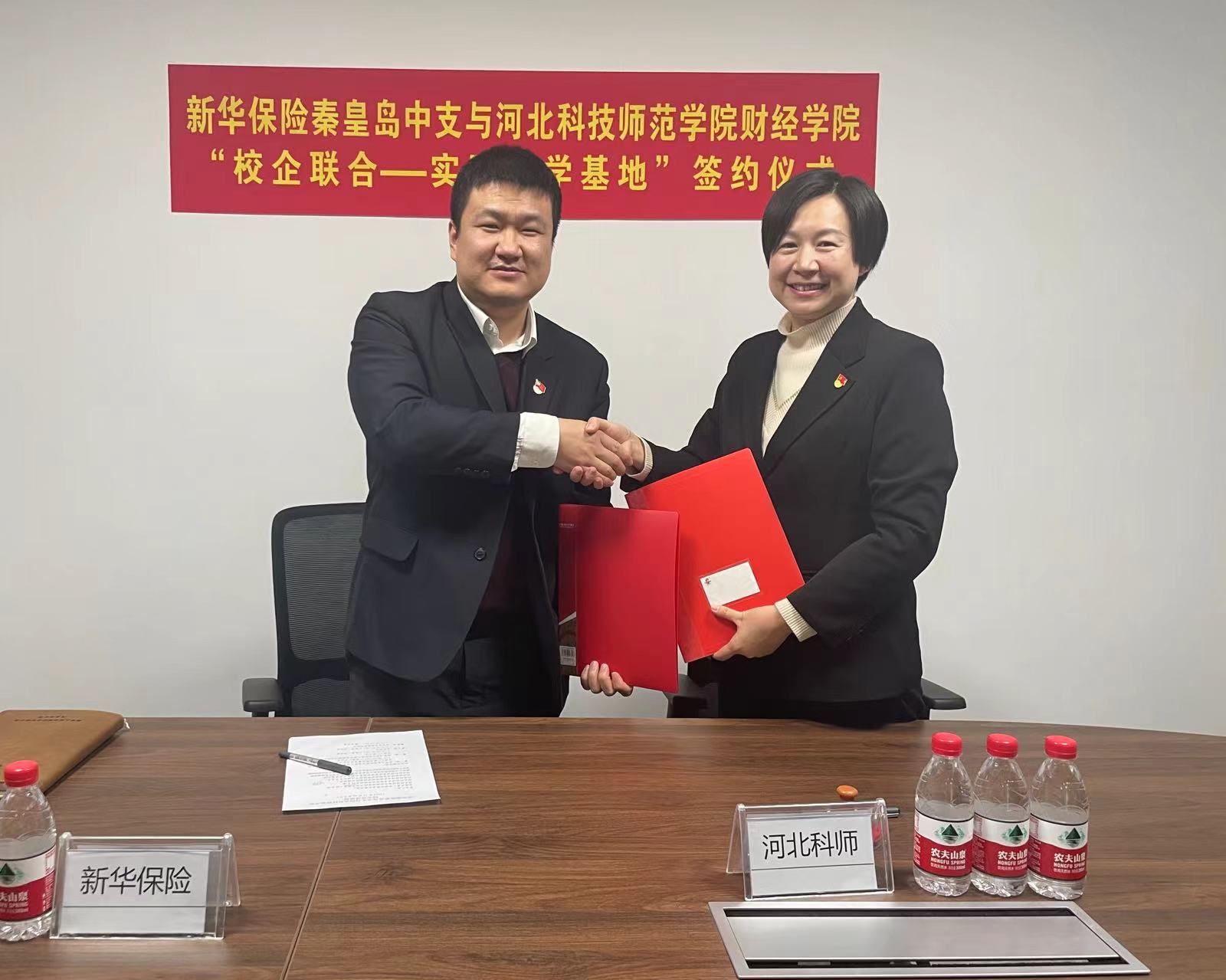
- 财达证券秦皇岛分公司走进我...06/14
- 学院召开考公经验分享会05/15
- 招生就业处到威人斯所有官方网站开展毕业生...05/10
- 学院开展证券行业发展与创新...04/19
- 学院开展“求职第一步,开启...04/19
- 学院召开毕业生就业工作推进会议03/20
- 学院开展“教育部24365招聘平...03/17
更多+图片展示
Copyright©2010 All Right Reserved 版权所有:威人斯所有官方网站 - 威尼斯会员登录
地址:中国河北省秦皇岛市河北大街西段360号 邮编:066004 联系电话:0335-8058250
领导信箱:jmxpgb@126.com